
The Importance of Influenza Diagnostic Surveillance and the Role of ATCC in Providing Reliable and Authenticated Analytical Reference Materials
Read the white paper to learn about the significance of influenza diagnostic surveillance and how ATCC is contributing to these efforts with authenticated analytical reference materials.
MoreATCC Cell Line Land: Advancing Transcriptome Sequencing for Cell Lines
Read the white paper to learn how ATCC Cell Line Land can help researchers confidently contribute to the advancement of scientific knowledge in a reliable and transparent manner, ensuring the integrity of their data and findings.
More White paper
White paper
Authentication of Prokaryotes at ATCC
Read this white paper to explore how prokaryotes are authenticated at ATCC.
More White paper
White paper
Bacteriophage Therapy An Alternative Approach for Treating Multidrug Resistant Infections
Read this white paper to learn more about multidrug resistance and how alternative treatment such as bacteriophage therapy may offer a promising mechanism for the eradication of infection.
More White paper
White paper
Bioluminescence Imaging of Live Animal Models
In this white paper we review the current models and techniques used in cancer research and discuss the use of bioluminescence imaging for the precise histological measurement of tumors.
More White paper
White paper
Challenges and solutions in the development and validation of molecular based assays
Discover more about the challenges associated with evaluating the analytical performance of a molecular assay and find guidance on selecting the ideal reference materials.
More White paper
White paper
CRISPRCas9 engineered cancer models the next step forward
ATCC has been steadily expanding an array of CRISPR/Cas9 genome-editing capabilities and leveraging its extensive library of human cell lines for the development of custom-engineered, biomarker-specific human cancer models.
More White paper
White paper
Drug Resistance in Cancer
This white paper examines how the next generation of advanced cancer models are vital tools for furthering our understanding drug-resistance mechanisms and will help lead to faster development cycles and lower failure rates for both anticancer drug development and evaluation of combination therapy strategies.
More White paper
White paper
Enhance the Development of Rapid Microbiological Methods
Read this white paper to learn how molecular-based rapid microbiological methods provide contemporary technologies used to quickly and sensitively detect, enumerate, and identify microorganisms.
More White paper
White paper
Improving accuracy and reproducibility in life science research
In this white paper, we review several predominant factors affecting reproducibility in life science research and outline efforts aimed at improving the situation.
More White paper
White paper
Infectious Disease Assay Development: Choosing the Appropriate External Controls
Here, we will discuss the importance of choosing the appropriate external controls, and will provide information on how to select the appropriate cultures and nucleic acids for your tests.
More White paper
White paper
Infectious disease assay development determining the limit of detection
In this white paper, we will discuss the importance of determining the detection limit in establishing analytical sensitivity, and will provide information on how to establish this parameter when evaluating your experimental design.
More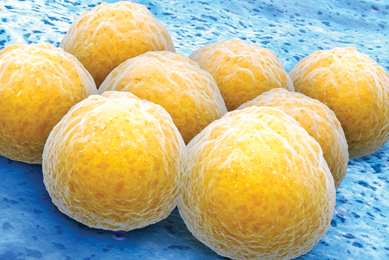 White paper
White paper
Infectious disease assay development establishing inclusivity exclusivity
In this white paper, we discuss the importance of inclusivity/exclusivity testing, and will provide information on how to establish these parameters when evaluating your experimental design.
More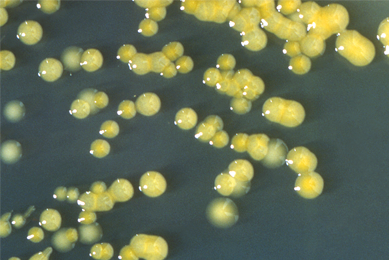 White paper
White paper
Microbiological Quality Control of Pharmaceutical Products
To ensure product safety, pharmaceutical companies must be versed in the important role of microbiological testing in product research and development, process validation, manufacturing, and quality control. Read this white paper to learn how ATCC supports the maintenance of product integrity, reputation, and safety by providing top-quality, fully characterized strains in a ready-to-use, familiar format.
More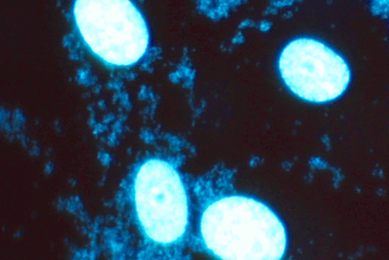 White paper
White paper
Mycoplasma quality control of cell substrates and biopharmaceuticals
In this article, we will discuss the effects of mycoplasma contamination, how this form of adulteration can affect cell-based drug development, and several quality control techniques and related products that can be used in the detection of mycoplasma contamination.
More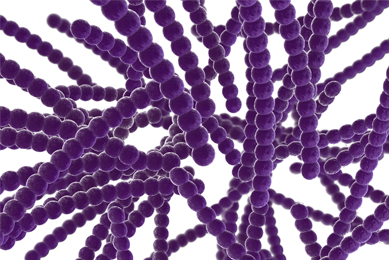 White paper
White paper
Pneumococcal Polysaccharides – Advancing Research and Development
In this white paper, we discuss the importance of capsular polysaccharides in Streptococcus pneumoniae infection and describe the use of purified serotype-specific polysaccharides in the development and evaluation of pneumococcal vaccines, the verification of novel immunoassays, and tracking bacterial disease epidemiology.
MoreReporter-Labeled Strains Powerful Tools for Microbiology Research
In this white paper, we highlight the reporters commonly used in microbiological research and provide detail on the various applications of reporter-labeled strains.
More White paper
White paper
Storing Biological Materials for Pharmaceutical R&D
Read this white paper for an overview of the relationship between the evolving practices of modern biorepositories and the current trends in pharmaceutical R&D.
More White paper
White paper
Synthetic Nucleic Acids for the Development and Evaluation of In Vitro Diagnostic Devices
In this white paper, we discuss the co-circulation and potential co-infection of the mosquito-borne Dengue (DENV), Chikungunya (CHIKV), and Zika (ZIKV) viruses; the challenges associated with the identification and discernment of these viruses; and the need for authenticated control materials for the validation of in vitro diagnostic tools.
More White paper
White paper
The Need for Standards in Human Microbiome Research
The human microbiome changes over time with respect to lifestyle, the environment, and exposure to disease. This individual biology is a powerful tool for personalized healthcare and precision medicine.
More White paper
White paper
The Need for Tuberculosis Reference Standards in Vaccine Development
In this white paper, we discuss how vaccine manufacturers can improve tuberculosis vaccine performance by utilizing a single, consensus biological reference standard that demonstrates high levels of protection when used in vaccine development.
More White paper
White paper
The Rise of Multidrug-Resistant Strains and Need for New Therapeutic Approaches
In this white paper, we discuss the growing threat of antimicrobial resistance, novel therapeutic approaches, and the importance of antimicrobial-resistant reference strains in reducing the emergence and spread of multidrug-resistant infections.
MoreView additional resources
Application Notes
Read our application notes for high-quality data exploring the development, validation, and application of ATCC products.
View Application Notes
Webinars
Watch our expanding collection of webinars to learn more about the innovative research and development being performed by thought leaders in science.
Watch the WebinarsCulture Guides
Download these useful guides and start with fresh authenticated cells and strains from ATCC to achieve the best results.
Read the Guides