Standards for next-generation sequencing
The complexities associated with 16S rRNA community profiling and shotgun metagenomics methods have made assay standardization challenging. ATCC has the solution: next-generation sequencing (NGS) standards. Throughout each stage of your workflow, NGS standards enable you to optimize your diverse research applications with confidence and improve the consistency and reproducibility of your data run after run. These standards support a broad array of applications ranging from method optimization to data interpretation, and they serve as superior controls for assay development and validation on any platform.
|
|
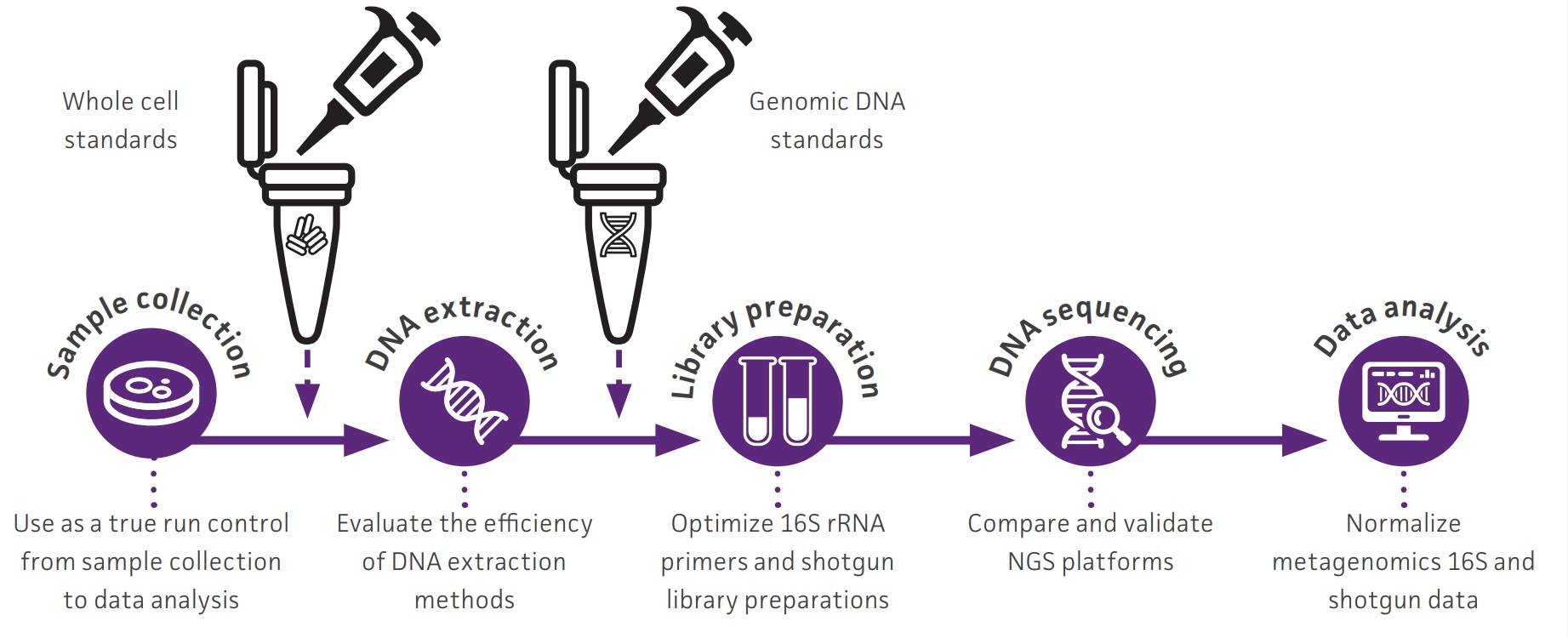
Explore how NGS Standards can optimize your research
Microbiome research
Although there is already a wealth of information on the human microbiome, much is still unknown. As analyses become more detailed, research will be challenged with burgeoning data sets and parsing information into meaningful insights through translational research. Discover how NGS Standards can help you optimize your metagenomic workflows and improve assay consistency.
Explore nowMolecular diagnostics
Molecular assays are powerful tools for the detection and identification of clinically relevant microbial pathogens. While these assays exhibit numerous benefits, they must be properly validated to guarantee accurate results and uncompromised data. Discover how NGS Standards can help you optimize your assay development process and improve the consistency of your data run after run.
Learn moreReference-quality sequences
Through the ATCC Genome Portal, you can easily search, access, and analyze thousands of reference-quality genome sequences. Our optimized methodology is designed to achieve complete, circularized (when biologically appropriate), and contiguous genomic elements by using short-read (virology collection) and hybrid (bacteriology, mycology, and protistology collections) assembly techniques. We then take our workflow one step further by accompanying each stage of the process with rigorous quality control analyses that ensure the highest quality data. Only the data that passes all quality control criteria are published to the ATCC Genome Portal. Visit the portal today to find the high-quality data you need for your research.
Visit the portalCheck out our data
 Application note
Application note
Development and Evaluation of Next-generation Sequencing Standards for Virome Research
In this study, we describe the development and quality control of the ATCC virome standards and present application data demonstrating their use throughout the different stages of a typical virome analysis workflow.
More Application note
Application note
Use of Recombinant Bacteria with Unique Tags as Spike-in Controls for Microbiome Studies
In this application note, we describe the construction and application of ATCC Spike-in Standards as a tool for quantitative metagenomic analysis.
More Poster
Poster
Authenticated Microbial Reference Genomes for Microbiome Analysis
This is a poster presented at World Microbe Forum 2021 that explores the generation of authenticated reference genomes for use in microbiome research workflows.
More